Our ECM Platform
It all starts with the patient
We have pioneered the world’s first patient-centric extracellular matrix-based discovery platform to identify and validate novel targets and biomarkers, and support the discovery and development of transformative therapies for fibrosis, inflammation and solid tumours.
Using our platform technology, we have the unique ability to use tissue- and disease-specific human ECM scaffolds to model and study the mechanisms driving disease biology. We deeply characterize both patient samples and models in terms of ECM composition and spatial biology and unite molecular with clinical and imaging data in our proprietary bioinformatics platform. This integrative approach ultimately leads us to targets and therapeutics which will benefit the patients who are at the centre of everything we do.

Introducing our cross-organ, cross-modality platform
Engitix’s platform is underpinned by an extensive bioarchive, one of the world’s largest ECM databases, the Engitomix proprietary bioinformatics platform, and best-in-class, human in vitro 3-D cell culture bioassays. These unique capabilities transform our ability to identify new targets and biomarkers, investigate novel mechanisms of action, and more accurately predict the efficacy of therapeutic candidates. Our platform is versatile and has been successfully adapted to support discovery efforts across a range of organs, indications, and therapeutic modalities.
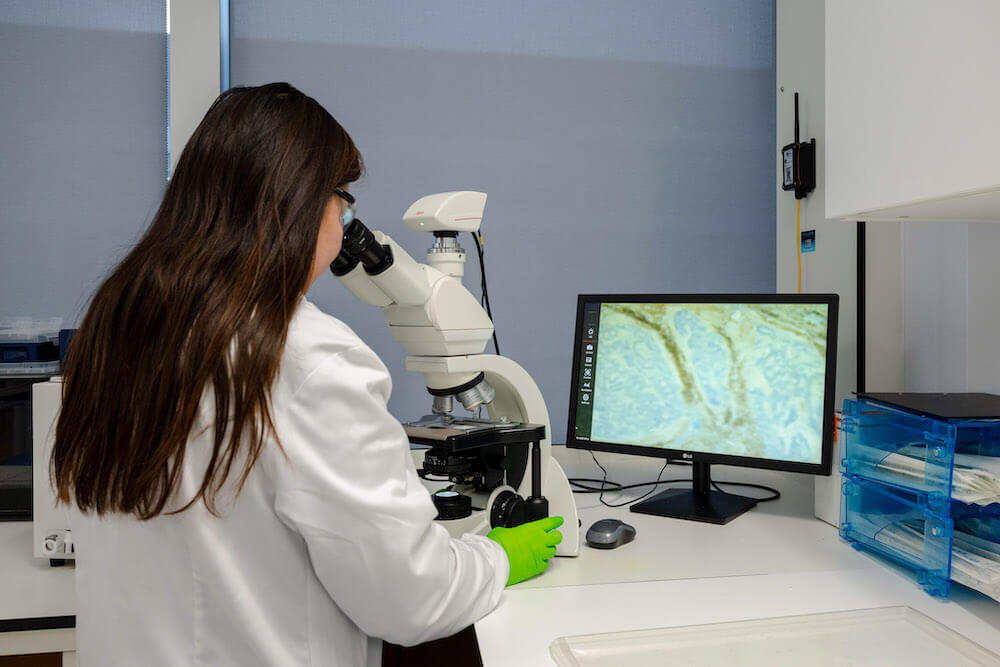
Biosamples and Biodata
We have our own biospecimens archive containing high-quality patient samples with associated clinical information, as well as from our strong network of partner hospitals, biobanks and KOLs worldwide.
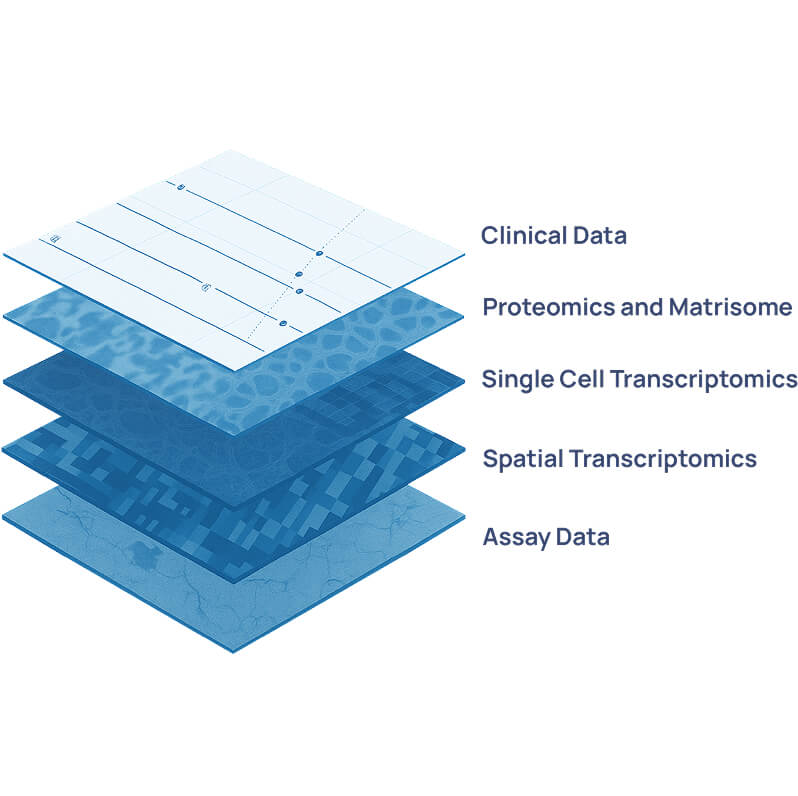
Engitomix Bioinformatics platform
The Engitomix Bioinformatics Platform integrates multi-omics and imaging data to decode the complexity of the extracellular matrix (ECM) and its impact on disease. Built around one of the world’s largest human ECM datasets, Engitomix unites mass-spectrometry, RNA-sequencing, and advanced imaging modalities to capture a holistic view of tissue- and disease-specific ECM biology.
Using our proprietary algorithms and AI/ML-driven analytics, Engitomix integrates these molecular profiles with well-curated clinical and patient outcome data. This enables the discovery of patterns and signatures that reveal how ECM composition and cell–matrix interactions evolve across disease stages and therapeutic responses.
By modelling these multidimensional datasets, Engitomix can prioritise novel targets and biomarkers with direct clinical relevance — accelerating the translation of ECM biology into precision therapies for fibrosis, inflammation, and solid tumours.

Human ECM Composition and Biology
We use our ECM platform to perform wet lab experiments in order to understand both the composition of the healthy and diseased acellular environment as well as the biology of this microenvironment in driving disease progression.
1. Decellularization
The composition of the disease-specific acellular ECM is characterised by proteomic analysis of decellularized tissues (‘matrisome’).
2. ECM solution to 3D model
Deconvoluting the biological impact of the ECM is evaluated by reseeding disease-relevant cell types into ECMs scaffolds or hydrogels.
Using these bioassays to assess molecular and functional changes in the cells, we can decode the changes to the ECM relevant for the disease of interest.


Translational 3D ECM screening platform
3D-ECM tissue-specific and disease-specific phenotypic screening.
Incorporating human ECM into our bioassays recreates a disease-relevant microenvironment to allow candidate mechanisms and drugs to be tested in a highly translational context. Engitix’s proprietary bioassays allow for a range of ECM-related biological activities and processes to be evaluated. These unique capabilities will enable differentiated therapeutic strategies to be discovered and transformational therapies to be developed. This patient-centric platform and discovery strategy has the potential to accelerate discovery, improve the accuracy of predicting candidate efficacy, and reduce the risk of failing to translate in clinical trials.


